Note
Go to the end to download the full example code.
Fitting NuSTAR Spectra: Glesener et al. 2020 comparison#
A real example from [Glesener2020] of fitting two NuSTAR spectra simultaneously with gain correction.
import warnings
import matplotlib.pyplot as plt
import numpy as np
from numpy.exceptions import VisibleDeprecationWarning
from parfive import Downloader
from sunkit_spex.legacy.fitting.fitter import Fitter
warnings.filterwarnings("ignore", category=RuntimeWarning)
try:
warnings.filterwarnings("ignore", category=VisibleDeprecationWarning)
except AttributeError:
warnings.filterwarnings("ignore", category=np.exceptions.VisibleDeprecationWarning)
Set up some plotting numbers
spec_single_plot_size = (8, 10)
spec_plot_size = (25, 10)
spec_font_size = 18
default_font_size = 10
x_limits, y_limits = [1.6, 8.5], [1e-1, 1e3]
# Let’s fit a more realistic example
Try fitting the spectra presented in [Glesener et al. 2020](https://iopscience.iop.org/article/10.3847/2041-8213/ab7341). The spectrum presented shows clear evidence of non-thermal emission in an A5.7 microflare.
## Let’s recreate Figure 4 (left) where NuSTAR FPMB is fitted with a thermal+cold thick target model.
set up plotting info stuff
gles_xlims, gles_ylims = [2, 12], [1e1, 1e4]
Download data files
dl = Downloader()
base_url = "https://sky.dias.ie/public.php/dav/files/BHW6y6aXiGGosM6/nustar/Glesener2020/"
file_names = [
"nu20312001001B06_cl_grade0_sr_grp.pha",
"nu20312001001B06_cl_grade0_sr.arf",
"nu20312001001B06_cl_grade0_sr.rmf",
"nu20312001001A06_cl_grade0_sr_grp.pha",
"nu20312001001A06_cl_grade0_sr.arf",
"nu20312001001A06_cl_grade0_sr.rmf",
]
for fname in file_names:
dl.enqueue_file(base_url + fname, path="../../nustar/Glesener2020/")
files = dl.download()
Files Downloaded: 0%| | 0/6 [00:00<?, ?file/s]
nu20312001001B06_cl_grade0_sr.arf: 0%| | 0.00/63.4k [00:00<?, ?B/s]
nu20312001001B06_cl_grade0_sr.rmf: 0%| | 0.00/27.0M [00:00<?, ?B/s]
nu20312001001B06_cl_grade0_sr_grp.pha: 0%| | 0.00/141k [00:00<?, ?B/s]
nu20312001001A06_cl_grade0_sr.arf: 0%| | 0.00/63.4k [00:00<?, ?B/s]
nu20312001001A06_cl_grade0_sr_grp.pha: 0%| | 0.00/141k [00:00<?, ?B/s]
nu20312001001B06_cl_grade0_sr.arf: 2%|▏ | 1.02k/63.4k [00:00<00:09, 6.64kB/s]
nu20312001001B06_cl_grade0_sr_grp.pha: 1%| | 1.02k/141k [00:00<00:21, 6.54kB/s]
nu20312001001A06_cl_grade0_sr.arf: 2%|▏ | 1.02k/63.4k [00:00<00:09, 6.57kB/s]
nu20312001001B06_cl_grade0_sr.rmf: 0%| | 1.02k/27.0M [00:00<1:13:07, 6.16kB/s]
nu20312001001A06_cl_grade0_sr_grp.pha: 1%| | 1.02k/141k [00:00<00:23, 5.98kB/s]
nu20312001001B06_cl_grade0_sr_grp.pha: 18%|█▊ | 25.6k/141k [00:00<00:00, 121kB/s]
nu20312001001A06_cl_grade0_sr.arf: 53%|█████▎ | 33.8k/63.4k [00:00<00:00, 160kB/s]
nu20312001001B06_cl_grade0_sr.arf: 66%|██████▋ | 42.0k/63.4k [00:00<00:00, 137kB/s]
Files Downloaded: 17%|█▋ | 1/6 [00:00<00:03, 1.39file/s]
nu20312001001B06_cl_grade0_sr_grp.pha: 65%|██████▌ | 92.2k/141k [00:00<00:00, 341kB/s]
nu20312001001B06_cl_grade0_sr.rmf: 0%| | 42.0k/27.0M [00:00<03:23, 132kB/s]
nu20312001001A06_cl_grade0_sr_grp.pha: 30%|██▉ | 42.0k/141k [00:00<00:00, 131kB/s]
nu20312001001B06_cl_grade0_sr_grp.pha: 94%|█████████▎| 132k/141k [00:00<00:00, 362kB/s]
Files Downloaded: 50%|█████ | 3/6 [00:00<00:00, 4.31file/s]
nu20312001001B06_cl_grade0_sr.rmf: 0%| | 116k/27.0M [00:00<01:23, 321kB/s]
nu20312001001A06_cl_grade0_sr_grp.pha: 89%|████████▉ | 126k/141k [00:00<00:00, 350kB/s]
nu20312001001A06_cl_grade0_sr.rmf: 0%| | 0.00/26.8M [00:00<?, ?B/s]
nu20312001001B06_cl_grade0_sr.rmf: 1%| | 247k/27.0M [00:00<00:42, 626kB/s]
nu20312001001A06_cl_grade0_sr.rmf: 0%| | 1.02k/26.8M [00:00<1:11:19, 6.26kB/s]
nu20312001001B06_cl_grade0_sr.rmf: 2%|▏ | 460k/27.0M [00:00<00:24, 1.09MB/s]
nu20312001001A06_cl_grade0_sr.rmf: 0%| | 99.3k/26.8M [00:00<00:57, 461kB/s]
nu20312001001B06_cl_grade0_sr.rmf: 4%|▎ | 983k/27.0M [00:00<00:11, 2.35MB/s]
nu20312001001A06_cl_grade0_sr.rmf: 1%| | 263k/26.8M [00:00<00:28, 929kB/s]
nu20312001001B06_cl_grade0_sr.rmf: 7%|▋ | 1.93M/27.0M [00:00<00:05, 4.51MB/s]
nu20312001001A06_cl_grade0_sr.rmf: 2%|▏ | 640k/26.8M [00:00<00:13, 1.96MB/s]
nu20312001001B06_cl_grade0_sr.rmf: 14%|█▍ | 3.89M/27.0M [00:00<00:02, 9.06MB/s]
nu20312001001A06_cl_grade0_sr.rmf: 5%|▌ | 1.40M/26.8M [00:00<00:06, 3.86MB/s]
nu20312001001B06_cl_grade0_sr.rmf: 28%|██▊ | 7.69M/27.0M [00:01<00:01, 17.3MB/s]
nu20312001001A06_cl_grade0_sr.rmf: 11%|█ | 2.90M/26.8M [00:00<00:03, 7.49MB/s]
nu20312001001B06_cl_grade0_sr.rmf: 41%|████ | 11.1M/27.0M [00:01<00:00, 20.8MB/s]
nu20312001001A06_cl_grade0_sr.rmf: 21%|██ | 5.50M/26.8M [00:00<00:01, 13.3MB/s]
nu20312001001B06_cl_grade0_sr.rmf: 55%|█████▍ | 14.8M/27.0M [00:01<00:00, 23.7MB/s]
nu20312001001A06_cl_grade0_sr.rmf: 33%|███▎ | 8.90M/26.8M [00:00<00:00, 19.6MB/s]
nu20312001001B06_cl_grade0_sr.rmf: 69%|██████▉ | 18.6M/27.0M [00:01<00:00, 25.4MB/s]
nu20312001001A06_cl_grade0_sr.rmf: 47%|████▋ | 12.6M/26.8M [00:01<00:00, 23.0MB/s]
nu20312001001B06_cl_grade0_sr.rmf: 83%|████████▎ | 22.3M/27.0M [00:01<00:00, 26.6MB/s]
nu20312001001A06_cl_grade0_sr.rmf: 61%|██████▏ | 16.4M/26.8M [00:01<00:00, 25.0MB/s]
nu20312001001B06_cl_grade0_sr.rmf: 97%|█████████▋| 26.1M/27.0M [00:01<00:00, 27.4MB/s]
Files Downloaded: 83%|████████▎ | 5/6 [00:02<00:00, 2.27file/s]
nu20312001001A06_cl_grade0_sr.rmf: 75%|███████▌ | 20.2M/26.8M [00:01<00:00, 26.3MB/s]
nu20312001001A06_cl_grade0_sr.rmf: 89%|████████▉ | 23.9M/26.8M [00:01<00:00, 27.3MB/s]
Files Downloaded: 100%|██████████| 6/6 [00:02<00:00, 2.66file/s]
Files Downloaded: 100%|██████████| 6/6 [00:02<00:00, 2.59file/s]
First, load in your data files, here we load in 2 spectra
_dir = "../../nustar/Glesener2020/"
# In the files here, the ARF and RMF file have different names to the PHA files so cannot use the PHA file name to help find the others so...
spec = Fitter(
pha_file=_dir + "nu20312001001B06_cl_grade0_sr_grp.pha",
arf_file=_dir + "nu20312001001B06_cl_grade0_sr.arf",
rmf_file=_dir + "nu20312001001B06_cl_grade0_sr.rmf",
)
Define model, here we go for a single isothermal model + cold thick model
spec.model = "f_vth + thick_fn"
Define fitting range
spec.energy_fitting_range = [2.8, 10.5]
Sort temperature param from f_vth
spec.params["T1_spectrum1"] = {"Value": 10.3, "Bounds": (1.1, 15)}
# emission measure param from f_vth
spec.params["EM1_spectrum1"] = {"Value": 0.5, "Bounds": (1e-2, 1e1)}
# electron flux param from thick_fn
spec.params["total_eflux1_spectrum1"] = {"Value": 2.1, "Bounds": (1e-3, 10)} # units 1e35 e^-/s
# electron index param from thick_fn
spec.params["index1_spectrum1"] = {"Value": 6.2, "Bounds": (3, 10)}
# electron low energy cut-off param from thick_fn
spec.params["e_c1_spectrum1"] = {"Value": 6.2, "Bounds": (1, 12)} # units keV
This fit requires altering the gain parameters
Gain parameters can be tweaked in the same way model parameters can.
The difference is that gain parameters all have specific starting values (slope=1, offset=0) and are frozen by default.
# from Glesener et al. 2020 which had a gain correction fixed at 0.95
spec.rParams["gain_slope_spectrum1"] = {"Status": "fixed", "Value": 0.95}
print(spec.rParams)
print(spec.show_rParams)
Status Value Bounds Error
gain_slope_spectrum1 frozen 0.95 (0.8, 1.2) (0.0, 0.0)
gain_offset_spectrum1 frozen 0.00 (-0.1, 0.1) (0.0, 0.0)
Param Status Value Bounds Error
(min, max) (-, +)
--------------------- ------ ---------- ----------- ------------------------
gain_slope_spectrum1 frozen 0.950 (0.8, 1.2) ( 0.00, 0.00)
gain_offset_spectrum1 frozen 0.000 (-0.1, 0.1) ( 0.00, 0.00)
Fit the model to the spectrum
spec.fit(tol=1e-8)
# Plot the result
plt.rcParams["font.size"] = spec_font_size
plt.figure(figsize=spec_single_plot_size)
axes, res_axes = spec.plot()
for a in axes:
a.set_xlim(gles_xlims)
a.set_ylim(gles_ylims)
plt.show()
plt.rcParams["font.size"] = default_font_size
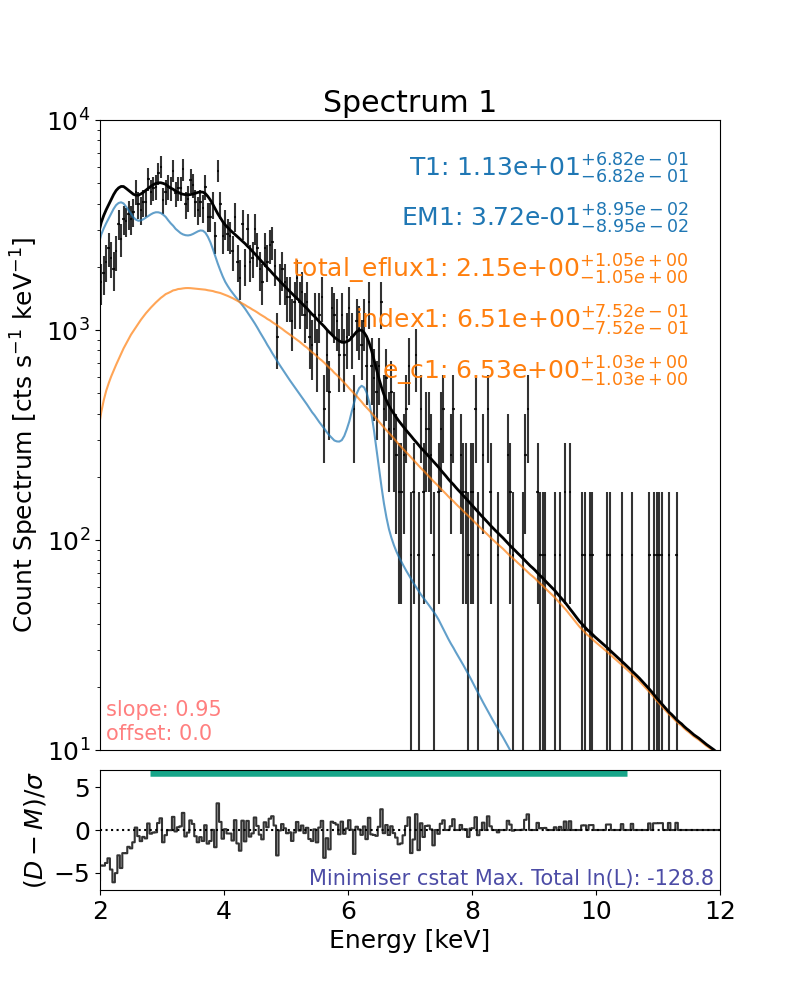
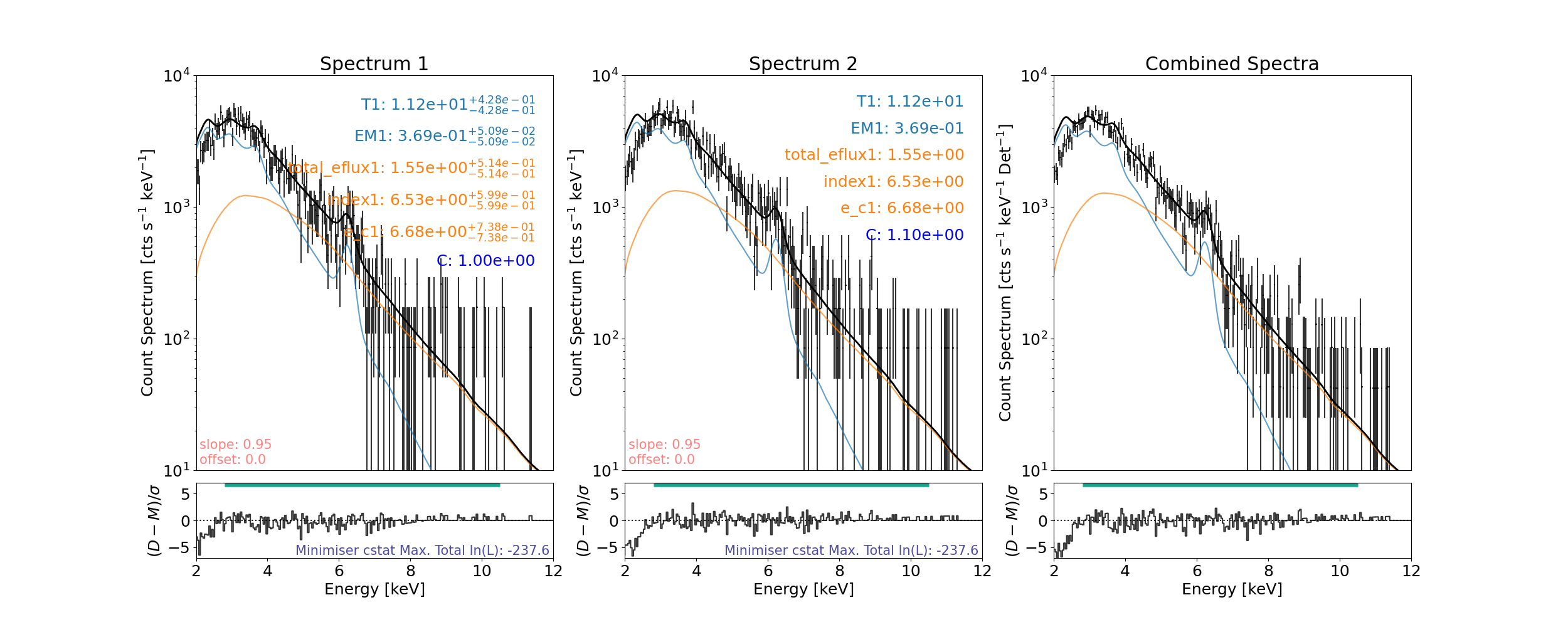
Let’s recreate something similar to Figure 3(c)
Both NuSTAR FPMs are fitted with a thermal+cold thick target model simultaneously.
In the original Figure 3(c), a broken power law is used as the cold thick target does not exist ins XPSEC.
First, load in your data files, here we load in 2 spectra
_dir = "../../nustar/Glesener2020/"
spec = Fitter(
pha_file=[_dir + "nu20312001001A06_cl_grade0_sr_grp.pha", _dir + "nu20312001001B06_cl_grade0_sr_grp.pha"],
arf_file=[_dir + "nu20312001001A06_cl_grade0_sr.arf", _dir + "nu20312001001B06_cl_grade0_sr.arf"],
rmf_file=[_dir + "nu20312001001A06_cl_grade0_sr.rmf", _dir + "nu20312001001B06_cl_grade0_sr.rmf"],
)
Define model, here we go for a single isothermal model + cold thick model
spec.model = "C*(f_vth + thick_fn)"
Define fitting range
spec.energy_fitting_range = [2.8, 10.5]
Sort temperature param from f_vth
spec.params["T1_spectrum1"] = {"Value": 10.3, "Bounds": (1.1, 15)}
# emission measure param from f_vth
spec.params["EM1_spectrum1"] = {"Value": 0.5, "Bounds": (1e-2, 1e1)}
# electron flux param from thick_fn
spec.params["total_eflux1_spectrum1"] = {"Value": 2.1, "Bounds": (1e-3, 10)} # units 1e35 e^-/s
# electron index param from thick_fn
spec.params["index1_spectrum1"] = {"Value": 6.2, "Bounds": (3, 10)}
# electron low energy cut-off param from thick_fn
spec.params["e_c1_spectrum1"] = {"Value": 6.2, "Bounds": (1, 12)} # units keV
# constant for systematic offset between FPMs, found to be about 1.1
spec.params["C_spectrum1"] = "frozen"
# constant for systematic offset between FPMs, found to be about 1.1
spec.params["C_spectrum2"] = {"Status": "fixed", "Value": 1.1}
# from Gles. 2020 which had a gain correction fixed at 0.95
spec.rParams["gain_slope_spectrum1"] = {"Status": "fixed", "Value": 0.95}
spec.rParams["gain_slope_spectrum2"] = spec.rParams["gain_slope_spectrum1"]
Fit the model to the spectrum
spec.fit(tol=1e-8)
# Plot the result
plt.rcParams["font.size"] = spec_font_size
plt.figure(figsize=spec_plot_size)
axes, res_axes = spec.plot()
for a in axes:
a.set_xlim(gles_xlims)
a.set_ylim(gles_ylims)
plt.show()
plt.rcParams["font.size"] = default_font_size
For the thermal and cold thick target total model we compare
Model Parameter |
OSPEX (Glesener et al. 2020) [*] |
This Work (just FPMB) |
This Work (FPMA&B) |
|---|---|---|---|
Temperature [MK] |
10.3\(\pm\)0.7 |
11.3\(\pm\)0.7 |
11.2\(\pm\)0.4 |
Emission Measure [cm-3] |
5.0 \(\pm\) 1.3\(\times\)10 45 |
3.7 \(\pm\) 0.9\(\times\)10 45 |
3.7 \(\pm\) 0.5\(\times\)10 45 |
Electron Flux [e$^{-}$ s$^{-1}$] |
2.1\(\pm\)1.1\(\times\)10 35 |
2.2\(\pm\)1.1\(\times\)10 35 |
1.6\(\pm\)0.5\(\times\)10 35 |
Index |
6.2\(\pm\)0.6 |
6.5\(\pm\)0.8 |
6.5\(\pm\)0.6 |
Low Energy Cut-off [keV] |
6.2\(\pm\)0.9 |
6.5\(\pm\)1.0 |
6.7\(\pm\)0.7 |
Now let’s recreate Figure 4 (right)
NuSTAR FPMB is fitted with a warm thick target model.
The warm thick target model helps to constrain the non-thermal emission with observed values (e.g., loop length, etc) and ties it to the thermal emission parameters.
First, load in your data files, here we load in 1 spectrum
define model, here we go for a single isothermal model + cold thick model
spec.model = "thick_warm"
define fitting range
spec.energy_fitting_range = [2.8, 10.5]
# electron flux param
spec.params["tot_eflux1_spectrum1"] = {"Value": 2, "Bounds": (1e-2, 10)}
# electron index param
spec.params["ind1_spectrum1"] = {"Value": 6, "Bounds": (3, 10)}
# electron low energy cut-off param
spec.params["ec1_spectrum1"] = {"Value": 7, "Bounds": (3, 12)}
# loop plasma temperature param
spec.params["loop_temp1_spectrum1"] = {"Value": 10, "Bounds": (5, 15)}
# plasma number density param
spec.params["plasma_d1_spectrum1"] = {"Value": 1, "Bounds": (1e-2, 1e1)} # units 1e10 cm^-3
# loop length param
spec.params["length1_spectrum1"] = {"Status": "fixed", "Value": 15} # units Mm
# from Gles. 2020 which had a gain correction fixed at 0.95
spec.rParams["gain_slope_spectrum1"] = {"Status": "fixed", "Value": 0.95}
fit the model to the spectrum
spec.fit(tol=1e-10)
# plot the result
plt.rcParams["font.size"] = spec_font_size
plt.figure(figsize=spec_single_plot_size)
axes, res_axes = spec.plot()
for a in axes:
a.set_xlim(gles_xlims)
a.set_ylim(gles_ylims)
plt.show()
plt.rcParams["font.size"] = default_font_size
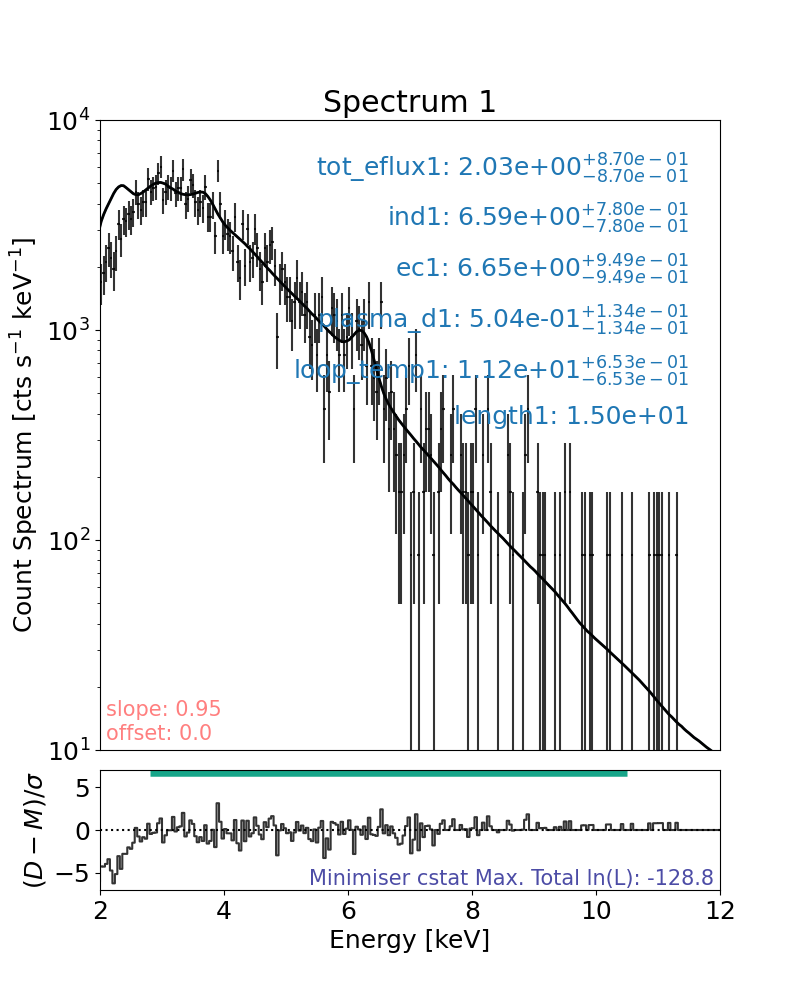
Minimum length>length
Fitting the warm thick target model to both FPMs simultaneously
First, load in your data files, here we load in 2 spectra
_dir = "../../nustar/Glesener2020/"
spec = Fitter(
pha_file=[_dir + "nu20312001001A06_cl_grade0_sr_grp.pha", _dir + "nu20312001001B06_cl_grade0_sr_grp.pha"],
arf_file=[_dir + "nu20312001001A06_cl_grade0_sr.arf", _dir + "nu20312001001B06_cl_grade0_sr.arf"],
rmf_file=[_dir + "nu20312001001A06_cl_grade0_sr.rmf", _dir + "nu20312001001B06_cl_grade0_sr.rmf"],
)
Define model, here we go for a single isothermal model + cold thick model
spec.model = "C*thick_warm"
Define fitting range
spec.energy_fitting_range = [2.8, 10.5]
# electron flux param
spec.params["tot_eflux1_spectrum1"] = {"Value": 2, "Bounds": (1e-3, 10)}
# electron index param
spec.params["ind1_spectrum1"] = {"Value": 6, "Bounds": (3, 10)}
# electron low energy cut-off param
spec.params["ec1_spectrum1"] = {"Value": 7, "Bounds": (1, 12)}
# loop plasma temperature param
spec.params["loop_temp1_spectrum1"] = {"Value": 10, "Bounds": (1.1, 15)}
# plasma number density param
spec.params["plasma_d1_spectrum1"] = {"Value": 1, "Bounds": (1e-2, 1e1)}
# loop length param
spec.params["length1_spectrum1"] = {"Status": "fixed", "Value": 15}
# constant for systematic offset between FPMs, found to be about 1.1
spec.params["C_spectrum1"] = "frozen"
spec.params["C_spectrum2"] = {"Status": "fixed", "Value": 1.1}
# from Gles. 2020 which had a gain correction fixed at 0.95
spec.rParams["gain_slope_spectrum1"] = {"Status": "fixed", "Value": 0.95}
spec.rParams["gain_slope_spectrum2"] = spec.rParams["gain_slope_spectrum1"]
Fit the model to the spectrum
spec.fit(tol=1e-5)
# Plot the result
plt.rcParams["font.size"] = spec_font_size
plt.figure(figsize=spec_plot_size)
axes, res_axes = spec.plot()
for a in axes:
a.set_xlim(gles_xlims)
a.set_ylim(gles_ylims)
plt.show()
plt.rcParams["font.size"] = default_font_size
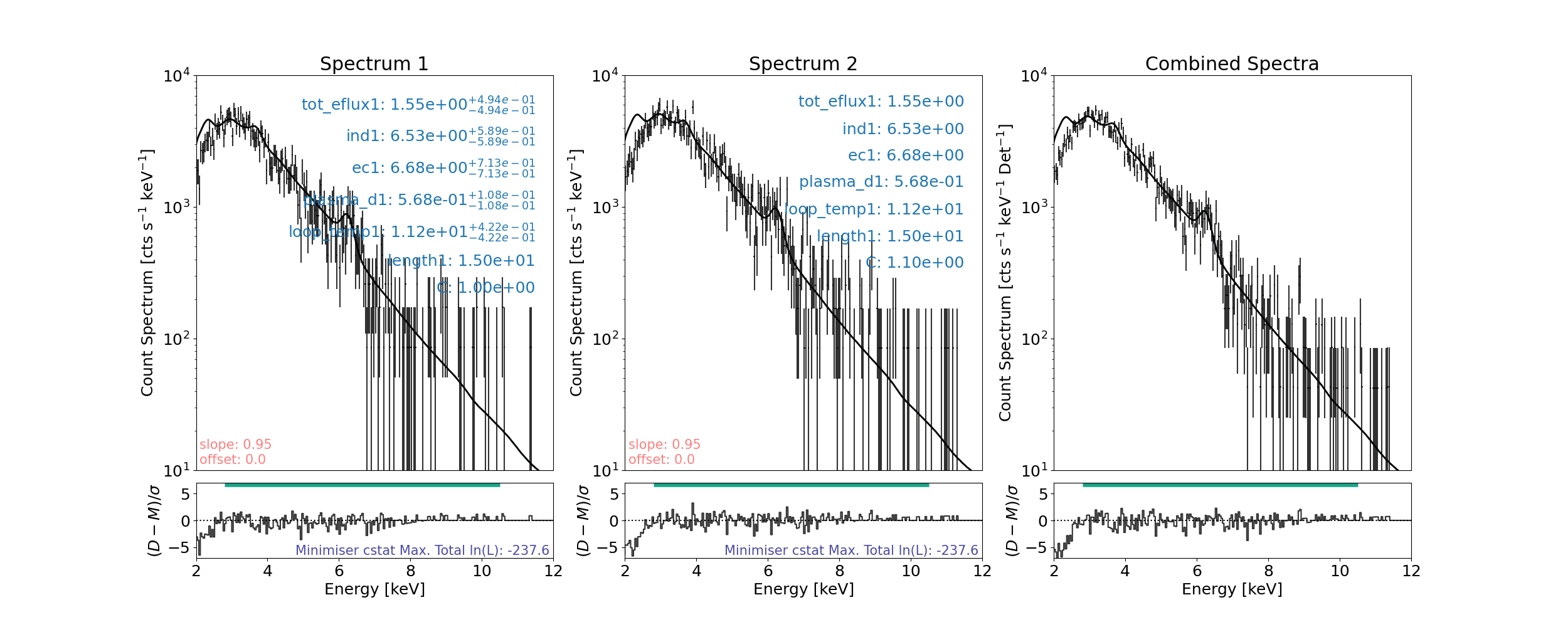
For warm thick target total model we compare
Model Parameter |
OSPEX (Glesener et al. 2020) |
This Work (just FPMB) |
This Work (FPMA&B) |
|---|---|---|---|
Temperature [MK] |
10.2\(\pm\)0.7 |
11.2\(\pm\)0.6 |
11.2\(\pm\)0.4 |
Plasma Density [cm-3] |
6.0 \(\pm\) 2.0\(\times\)10 9 |
5.0 \(\pm\) 1.3\(\times\)10 9 |
5.7 \(\pm\) 1.1\(\times\)10 9 |
Electron Flux [e$^{-}$ s$^{-1}$] |
1.8\(\pm\)0.8\(\times\)10 35 |
2.0\(\pm\)0.8\(\times\)10 35 |
1.6\(\pm\)0.5\(\times\)10 35 |
Index |
6.3\(\pm\)0.7 |
6.6\(\pm\)0.8 |
6.5\(\pm\)0.6 |
Low Energy Cut-off [keV] |
6.5\(\pm\)0.9 |
6.7\(\pm\)0.9 |
6.7\(\pm\)0.7 |
All parameter values appear to be within error margins (or extremely close). This is more impressive when the errors calculated in this work for the minimised values assumes the parameter’s have a Gaussian and independent posterior distribution (which is clearly not the case) and so these errors are likely to be larger; to be investigated with an MCMC.
The simultaneous fit of FPMA&B with the cold thick target model and the warm thick model is not able to be performed in OSPEX.
Total running time of the script: (2 minutes 16.829 seconds)