Note
Go to the end to download the full example code.
Fitting NuSTAR Spectra: Duncan et al. 2021 comparison#
This spectrum corresponds to the may1618 microflare in [Duncan2021].
An example of fitting multiple spectra simultaneously with 2 models where each model is allowed to vary at different times
We also allow the gain slope response parameter to vary.
import warnings
import matplotlib.pyplot as plt
import numpy as np
from numpy.exceptions import VisibleDeprecationWarning
from parfive import Downloader
from sunkit_spex.legacy.fitting.fitter import Fitter
warnings.filterwarnings("ignore", category=RuntimeWarning)
try:
warnings.filterwarnings("ignore", category=VisibleDeprecationWarning)
except AttributeError:
warnings.filterwarnings("ignore", category=np.exceptions.VisibleDeprecationWarning)
Set up some plotting numbers
spec_single_plot_size = (8, 10)
spec_plot_size = (25, 10)
spec_font_size = 18
default_font_size = 10
x_limits, y_limits = [1.6, 8.5], [1e-1, 1e3]
# set up plotting info stuff
dunc_xlims, dunc_ylims = [2.5, 11], [1e0, 4e4]
Download the example data
dl = Downloader()
base_url = "https://sky.dias.ie/public.php/dav/files/BHW6y6aXiGGosM6/nustar/Duncan2021/"
file_names = [
"nu80410201001A06_1618_p_chu2_N_sr.pha",
"nu80410201001A06_1618_p_chu2_N_sr.arf",
"nu80410201001A06_1618_p_chu2_N_sr.rmf",
"nu80410201001B06_1618_p_chu2_N_sr.pha",
"nu80410201001B06_1618_p_chu2_N_sr.arf",
"nu80410201001B06_1618_p_chu2_N_sr.rmf",
]
for fname in file_names:
dl.enqueue_file(base_url + fname, path="../../nustar/Duncan2021/")
files = dl.download()
Files Downloaded: 0%| | 0/6 [00:00<?, ?file/s]
nu80410201001B06_1618_p_chu2_N_sr.pha: 0%| | 0.00/153k [00:00<?, ?B/s]
nu80410201001A06_1618_p_chu2_N_sr.arf: 0%| | 0.00/63.4k [00:00<?, ?B/s]
nu80410201001B06_1618_p_chu2_N_sr.arf: 0%| | 0.00/63.4k [00:00<?, ?B/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 0%| | 0.00/30.5M [00:00<?, ?B/s]
nu80410201001A06_1618_p_chu2_N_sr.pha: 0%| | 0.00/153k [00:00<?, ?B/s]
nu80410201001B06_1618_p_chu2_N_sr.pha: 1%| | 1.02k/153k [00:00<00:23, 6.54kB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 0%| | 1.02k/30.5M [00:00<1:16:03, 6.68kB/s]
nu80410201001A06_1618_p_chu2_N_sr.arf: 2%|▏ | 1.02k/63.4k [00:00<00:09, 6.35kB/s]
nu80410201001B06_1618_p_chu2_N_sr.arf: 2%|▏ | 1.02k/63.4k [00:00<00:09, 6.30kB/s]
nu80410201001A06_1618_p_chu2_N_sr.pha: 1%| | 1.02k/153k [00:00<00:23, 6.51kB/s]
nu80410201001A06_1618_p_chu2_N_sr.pha: 17%|█▋ | 25.6k/153k [00:00<00:01, 120kB/s]
nu80410201001B06_1618_p_chu2_N_sr.pha: 28%|██▊ | 42.0k/153k [00:00<00:00, 137kB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 0%| | 42.0k/30.5M [00:00<03:43, 136kB/s]
nu80410201001A06_1618_p_chu2_N_sr.arf: 66%|██████▋ | 42.0k/63.4k [00:00<00:00, 135kB/s]
Files Downloaded: 17%|█▋ | 1/6 [00:00<00:04, 1.09file/s]
nu80410201001B06_1618_p_chu2_N_sr.arf: 66%|██████▋ | 42.0k/63.4k [00:00<00:00, 133kB/s]
nu80410201001A06_1618_p_chu2_N_sr.pha: 52%|█████▏ | 79.9k/153k [00:00<00:00, 291kB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 0%| | 108k/30.5M [00:00<01:40, 301kB/s]
nu80410201001A06_1618_p_chu2_N_sr.pha: 92%|█████████▏| 140k/153k [00:00<00:00, 406kB/s]
Files Downloaded: 67%|██████▋ | 4/6 [00:01<00:00, 4.91file/s]
nu80410201001B06_1618_p_chu2_N_sr.rmf: 0%| | 0.00/27.0M [00:00<?, ?B/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 1%| | 230k/30.5M [00:00<00:51, 591kB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 2%|▏ | 476k/30.5M [00:00<00:25, 1.17MB/s]
nu80410201001B06_1618_p_chu2_N_sr.rmf: 0%| | 1.02k/27.0M [00:00<1:15:44, 5.94kB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 3%|▎ | 984k/30.5M [00:00<00:12, 2.36MB/s]
nu80410201001B06_1618_p_chu2_N_sr.rmf: 0%| | 50.2k/27.0M [00:00<01:59, 226kB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 6%|▌ | 1.90M/30.5M [00:00<00:06, 4.43MB/s]
nu80410201001B06_1618_p_chu2_N_sr.rmf: 1%| | 140k/27.0M [00:00<00:54, 489kB/s]
nu80410201001B06_1618_p_chu2_N_sr.rmf: 1%|▏ | 345k/27.0M [00:00<00:25, 1.05MB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 11%|█▏ | 3.49M/30.5M [00:00<00:03, 7.00MB/s]
nu80410201001B06_1618_p_chu2_N_sr.rmf: 3%|▎ | 763k/27.0M [00:00<00:12, 2.11MB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 25%|██▍ | 7.57M/30.5M [00:01<00:01, 14.0MB/s]
nu80410201001B06_1618_p_chu2_N_sr.rmf: 6%|▌ | 1.57M/27.0M [00:00<00:06, 4.06MB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 33%|███▎ | 9.93M/30.5M [00:01<00:01, 16.4MB/s]
nu80410201001B06_1618_p_chu2_N_sr.rmf: 10%|█ | 2.76M/27.0M [00:00<00:03, 6.54MB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 41%|████ | 12.5M/30.5M [00:01<00:00, 18.7MB/s]
nu80410201001B06_1618_p_chu2_N_sr.rmf: 20%|█▉ | 5.29M/27.0M [00:00<00:01, 12.4MB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 49%|████▉ | 15.0M/30.5M [00:01<00:00, 20.5MB/s]
nu80410201001B06_1618_p_chu2_N_sr.rmf: 33%|███▎ | 8.82M/27.0M [00:00<00:00, 19.3MB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 57%|█████▋ | 17.5M/30.5M [00:01<00:00, 21.8MB/s]
nu80410201001B06_1618_p_chu2_N_sr.rmf: 47%|████▋ | 12.6M/27.0M [00:01<00:00, 22.8MB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 66%|██████▌ | 20.1M/30.5M [00:01<00:00, 22.9MB/s]
nu80410201001B06_1618_p_chu2_N_sr.rmf: 60%|██████ | 16.3M/27.0M [00:01<00:00, 25.2MB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 75%|███████▍ | 22.7M/30.5M [00:01<00:00, 23.9MB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 83%|████████▎ | 25.4M/30.5M [00:01<00:00, 24.6MB/s]
nu80410201001B06_1618_p_chu2_N_sr.rmf: 74%|███████▍ | 20.1M/27.0M [00:01<00:00, 26.6MB/s]
nu80410201001A06_1618_p_chu2_N_sr.rmf: 92%|█████████▏| 28.1M/30.5M [00:01<00:00, 25.1MB/s]
nu80410201001B06_1618_p_chu2_N_sr.rmf: 88%|████████▊ | 23.8M/27.0M [00:01<00:00, 27.5MB/s]
Files Downloaded: 100%|██████████| 6/6 [00:02<00:00, 2.16file/s]
Files Downloaded: 100%|██████████| 6/6 [00:02<00:00, 2.29file/s]
First, load in your data files, here we load in 2 spectra
Define model, here we go for 2 isothermal models
spec.model = "C*(f_vth + f_vth)"
Check the parameter table
print(spec.params)
Status Value Bounds Error
T1_spectrum1 free 1.0 (0.0, None) (0.0, 0.0)
EM1_spectrum1 free 1.0 (0.0, None) (0.0, 0.0)
T2_spectrum1 free 1.0 (0.0, None) (0.0, 0.0)
EM2_spectrum1 free 1.0 (0.0, None) (0.0, 0.0)
C_spectrum1 free 1.0 (0.0, None) (0.0, 0.0)
T1_spectrum2 tie_T1_spectrum1 1.0 (0.0, None) (0.0, 0.0)
EM1_spectrum2 tie_EM1_spectrum1 1.0 (0.0, None) (0.0, 0.0)
T2_spectrum2 tie_T2_spectrum1 1.0 (0.0, None) (0.0, 0.0)
EM2_spectrum2 tie_EM2_spectrum1 1.0 (0.0, None) (0.0, 0.0)
C_spectrum2 tie_C_spectrum1 1.0 (0.0, None) (0.0, 0.0)
freeze the ones we don’t want to vary
spec.params["C_spectrum1"] = {"Status": "frozen"}
Set initial values
spec.params["T1_spectrum1"] = {"Value": 4.1, "Bounds": (2.5, 6)}
spec.params["EM1_spectrum1"] = {"Value": 14, "Bounds": (1e0, 1e2)}
spec.params["T2_spectrum1"] = {"Value": 10, "Bounds": (5, 15)}
spec.params["EM2_spectrum1"] = {"Value": 0.46, "Bounds": (1e-4, 10)}
spec.params["C_spectrum2"] = {"Status": "free", "Bounds": (0.5, 2)}
Fit lower energy range with the first thermal model first
spec.params["T2_spectrum1"] = "frozen"
spec.params["EM2_spectrum1"] = "frozen"
spec.energy_fitting_range = [2.5, 4]
spec.fit(tol=1e-6)
print(spec.params)
print(spec.rParams)
Status ... Error
T1_spectrum1 free ... (0.05064320644296751, 0.05064320644296751)
EM1_spectrum1 free ... (0.11786784508763132, 0.11786784508763132)
T2_spectrum1 frozen ... (0.0, 0.0)
EM2_spectrum1 frozen ... (0.0, 0.0)
C_spectrum1 frozen ... (0.0, 0.0)
T1_spectrum2 tie_T1_spectrum1 ... (0.0, 0.0)
EM1_spectrum2 tie_EM1_spectrum1 ... (0.0, 0.0)
T2_spectrum2 tie_T2_spectrum1 ... (0.0, 0.0)
EM2_spectrum2 tie_EM2_spectrum1 ... (0.0, 0.0)
C_spectrum2 free ... (0.005591092789415897, 0.005591092789415897)
[10 rows x 4 columns]
Status Value Bounds Error
gain_slope_spectrum1 frozen 1.0 (0.8, 1.2) (0.0, 0.0)
gain_offset_spectrum1 frozen 0.0 (-0.1, 0.1) (0.0, 0.0)
gain_slope_spectrum2 frozen 1.0 (0.8, 1.2) (0.0, 0.0)
gain_offset_spectrum2 frozen 0.0 (-0.1, 0.1) (0.0, 0.0)
Now fit higher energy range with the second thermal model
spec.params["T1_spectrum1"] = "frozen"
spec.params["EM1_spectrum1"] = "frozen"
spec.params["C_spectrum2"] = "frozen"
spec.params["T2_spectrum1"] = "free"
spec.params["EM2_spectrum1"] = "free"
Need the gain slope to vary too for this microflare but only needed for the 6.7 keV line
print(spec.rParams)
spec.rParams["gain_slope_spectrum1"] = "free"
spec.rParams["gain_slope_spectrum2"] = spec.rParams["gain_slope_spectrum1"]
spec.energy_fitting_range = [4, 10.8]
spec.fit(tol=1e-6)
print(spec.params)
print(spec.rParams)
Status Value Bounds Error
gain_slope_spectrum1 frozen 1.0 (0.8, 1.2) (0.0, 0.0)
gain_offset_spectrum1 frozen 0.0 (-0.1, 0.1) (0.0, 0.0)
gain_slope_spectrum2 frozen 1.0 (0.8, 1.2) (0.0, 0.0)
gain_offset_spectrum2 frozen 0.0 (-0.1, 0.1) (0.0, 0.0)
Status ... Error
T1_spectrum1 frozen ... (0.05064320644296751, 0.05064320644296751)
EM1_spectrum1 frozen ... (0.11786784508763132, 0.11786784508763132)
T2_spectrum1 free ... (0.04480653528434264, 0.04480653528434264)
EM2_spectrum1 free ... (0.009298437380353358, 0.009298437380353358)
C_spectrum1 frozen ... (0.0, 0.0)
T1_spectrum2 tie_T1_spectrum1 ... (0.0, 0.0)
EM1_spectrum2 tie_EM1_spectrum1 ... (0.0, 0.0)
T2_spectrum2 tie_T2_spectrum1 ... (0.0, 0.0)
EM2_spectrum2 tie_EM2_spectrum1 ... (0.0, 0.0)
C_spectrum2 frozen ... (0.005591092789415897, 0.005591092789415897)
[10 rows x 4 columns]
Status ... Error
gain_slope_spectrum1 free ... (0.00159648836679688, 0.00159648836679688)
gain_offset_spectrum1 frozen ... (0.0, 0.0)
gain_slope_spectrum2 tie_gain_slope_spectrum1 ... (0.0, 0.0)
gain_offset_spectrum2 frozen ... (0.0, 0.0)
[4 rows x 4 columns]
Now free everything over full range
spec.params["T1_spectrum1"] = "free"
spec.params["EM1_spectrum1"] = "free"
spec.params["C_spectrum2"] = "free"
spec.energy_fitting_range = [3, 10.8]
spec.fit(tol=1e-10)
print(spec.params)
print(spec.rParams)
Status ... Error
T1_spectrum1 free ... (0.28729959370082825, 0.28729959370082825)
EM1_spectrum1 free ... (2.435618930205077, 2.435618930205077)
T2_spectrum1 free ... (0.09072453923762787, 0.09072453923762787)
EM2_spectrum1 free ... (0.027840404020151744, 0.027840404020151744)
C_spectrum1 frozen ... (0.0, 0.0)
T1_spectrum2 tie_T1_spectrum1 ... (0.0, 0.0)
EM1_spectrum2 tie_EM1_spectrum1 ... (0.0, 0.0)
T2_spectrum2 tie_T2_spectrum1 ... (0.0, 0.0)
EM2_spectrum2 tie_EM2_spectrum1 ... (0.0, 0.0)
C_spectrum2 free ... (0.006381199192249198, 0.006381199192249198)
[10 rows x 4 columns]
Status ... Error
gain_slope_spectrum1 free ... (0.0014050421379300571, 0.0014050421379300571)
gain_offset_spectrum1 frozen ... (0.0, 0.0)
gain_slope_spectrum2 tie_gain_slope_spectrum1 ... (0.0, 0.0)
gain_offset_spectrum2 frozen ... (0.0, 0.0)
[4 rows x 4 columns]
Plot the result
plt.rcParams["font.size"] = spec_font_size
plt.figure(figsize=spec_plot_size)
axes, res_axes = spec.plot()
for a in axes:
a.set_xlim(dunc_xlims)
a.set_ylim(dunc_ylims)
plt.show()
plt.rcParams["font.size"] = default_font_size
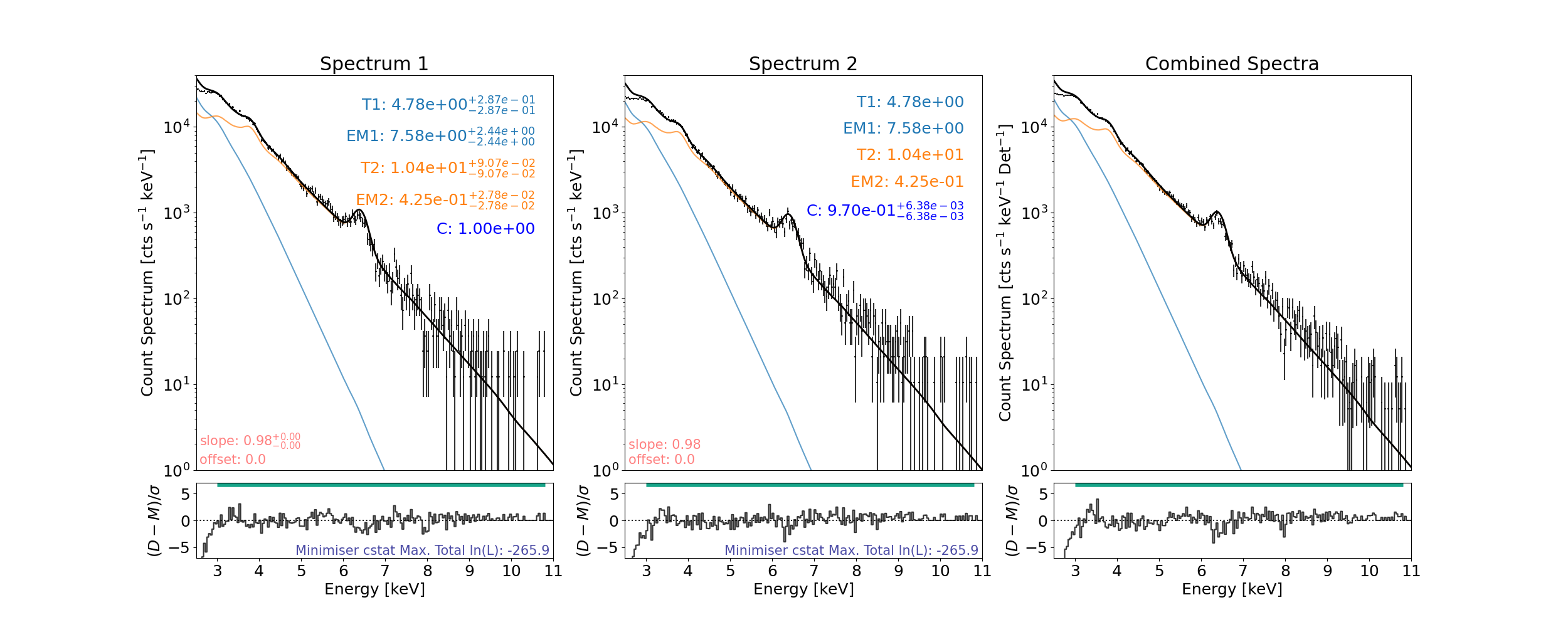
For the 2 thermal model fitting
Model Parameter |
XSPEC (Duncan et al. 2021) [*] |
This Work |
|---|---|---|
Temperature 1 [MK] |
\(4.1^{+0.1}_{-0.1}\) |
4.8\(\pm\)0.3 |
Emission Measure 1 [cm-3] |
\(1.4^{+0.6}_{-0.4}\times10^{47}\) |
7.6 \(\pm\) 2.4\(\times\)10 46 |
Temperature 2 [MK] |
\(10.00^{+0.03}_{-0.03}\) |
10.4\(\pm\)0.1 |
Emission Measure 2 [cm-3] |
\(4.6^{+0.1}_{-0.2}\times10^{45}\) |
4.3 \(\pm\) 0.3\(\times\)10 45 |
For the gain parameters
Model Parameter |
XSPEC (Duncan et al. 2021) |
This Work |
|---|---|---|
Gain Slope |
0.977\(\pm\)0.002 |
0.978\(\pm\)0.001 |
Although these values are slightly different, it is important to note that XSPEC and sunkit-spex work from different atomic databases. We also note that for a similar isothermal fit the temperature can drop/rise if the emission measure rises/drops and so fitting not just one but two of these models allows for these to vary more. We do see that this work (for this microflare) produces higher temperatures but correspondingly lower emission measures.
Total running time of the script: (0 minutes 36.948 seconds)